400-998-5282
专注多肽 服务科研
400-998-5282
专注多肽 服务科研
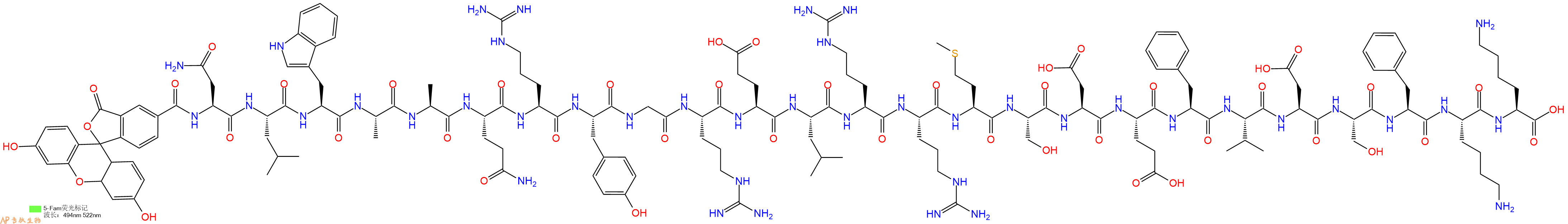
带 FAM 标签的人 BAD (103-127) (HY-P2468)。BAD (103-127) (human) 是 25-mer Bad 多肽,来自 BAD 的 BH3 结构域,具有拮抗 Bcl-xL 的作用。
编号:200662
CAS号:
单字母:5FAM-NLWAAQRYGRELRRMSDEFVDSFKK-OH
BAD (103-127) (human), FAM-labeled 是带 FAM 标签的人 BAD (103-127) (HY-P2468)。BAD (103-127) (human) 是 25-mer Bad 多肽,来自 BAD 的 BH3 结构域,具有拮抗 Bcl-xL 的作用。
BAD (103-127) (human), FAM-labeled is a FAM-labeled human BAD (103-127) (HY-P2468). BAD (103-127) (human), the 25-mer Bad peptide, is derived from the BH3 domain of BAD, can antagonize the function of Bcl-xL[1].
Definition
BAD is a member of the BCl-2 family of proteins and acts to promote apoptosis by forming heterodimers with the survival proteins BCL-2 and BCLXL, thus preventing them from binding with BAX1.
Discovery
BAD was originally identified in a yeast two hybrid system that was used to screen for BCL-2 interacting proteins1.
Classification
BAD belongs to the BH3 sub-family of proteins that also includes other pro-apoptotic proteins; BH3 BID, BIK, BLK, HRK, BNIP3 and BIML2.
Structural Characteristics
BAD peptides contain a highly conserved alpha-helical BH3 domain through which they form heterodimers with BCL-2 and BCLXL3. The BH3 domain is structurally defined as four-turn amphipathic a-helices, containing the sequence motif: Hy-X-X-X-Hy-X-X-X-Sm-D/E-X-Hy4. This domain is sufficient for pro-apoptotic functions of BAD.
Mode of action
BAD peptides are located on the outer-mitochondrail membrane. Activation of NGF or IL-3 receptors on the mitochondrial membrane mediates the activation of AKT or PKA holoenzyme respectively that result in the phosphorylation of BAD at Ser-136 and 1122,5. Phosphorylated BAD is translocated to the cytosol by phosphoserine binding protein. Following a death signal BAD is dephosporylated and found in association with BCL-XL-BCL-2 in which form BAD can exert its functions6.
Functions
BCL-2 family proteins that includes BAD play a pivotal role in deciding whether a cell will live or die by apoptosis. Pro-apoptotic function of BAD is triggered by growth factor deprivation in the cell that results in its dephosphorylation and activation7. Activated BAD binds to BCL-2-BCL-XL and releases BAK and BAX that initiate apoptosis. Increased BAD protein levels have been found in diseases like myocardial ischemia-reperfusion7. BAD is also implicated in cancer. In mouse models it has been found that decrease in BAD levels leads to malignancy8. Interestingly recent studies have shown that the tumor suppressor protein, p53 binds to BAD in response to DNA damage and in turn BAD triggers apoptosis of such cells thus maintaining cell physiology9.
References
1. Elizabeth Y, Jiping Z, Jennifer J, Boise LH, Craig B, Thompson and Stanley JK, (1995). Bad, a heterodimeric partner for Bcl-xL and Bcl-2, displaces bax and promotes cell death. Cell, 80, Issue 2, 285-91.
2. Atan G, James M.M, Stanley J.K (1999). BCL-2 family members and the mitochondria in apoptosis. Genes and Development, 13; 1899-1911.
3. Sabine O, Jose-L D, William H, Julia C, Yan W, Gary W, Steve C, Suzanne W, Lawrence CF, and Tilman O (1997). Structural properties of Human BAD. J Biol. Chem., 372, 49, 30866-892.
4. Beth L, Sangita S and Guido K (2008). Bcl-2 family members; Dual regulators of apoptosis and autophagy. Autophagy, 4:5, 600-606.
5. Zha J, Harada H, Yang E, Jockel J, Kormeyer SJ (1996c). Serine phosphorylation of death agonist BAD in response to survival factor results in binding to 14-3-3 not BCL-X(L). Cell, 87:619–628.
6. Zha J, Harada H, Osipov K, Jockel J, Waksman G, Korsmeyer SJ (1997). BH3 domain of BAD is required for heterodimerization with BCL-XL and pro-apoptotic activity. J. Biol. Chem., 272:24101–24104.
7. Åsa BG and Roberta AG (2007). Bcl-2 family members and apoptosis, taken to heart. Am J Physiol Cell Physiol., 292:45-51.
8. Zinkel S, Gross A and Yang E (2006). BCL2 family in DNA damage and cell cycle control. Cell Death and Differentiation, 13, 1351–1359.
9. Peng J, Wenjing D and Mian W (2007). p53 and BAD: Remote strangers become close friends, Cell Research, 17: 283–285.
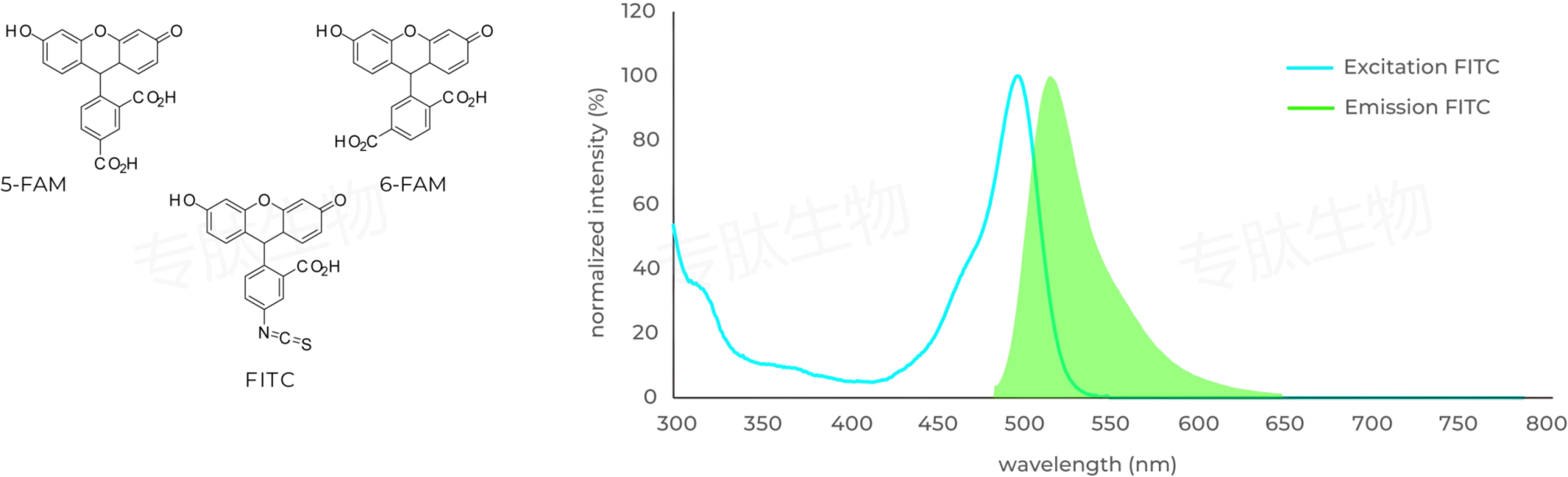
FAM标记说明:
Carboxyfluorescein (FAM) is fluorophor with an excitation at 492 nm ▉ and emission of 517 nm ▉. Donors like FAM and 5FAM are often paired together with acceptors (CPQ2) for FRET experiments.
FAM标记肽的相关文献:
Porous Silicon Nanoparticle Delivery of Tandem Peptide Anti-Infectives for the Treatment of Pseudomonas aeruginosa Lung Infections.
Kwon, Ester J., et al. Advanced Materials 29.35 (2017).
Ultrasensitive tumor-penetrating nanosensors of protease activity.
Kwon, Ester J., Jaideep S. Dudani, and Sangeeta N. Bhatia. Nature Biomedical Engineering 1 (2017): 0054.
Seneca Valley Virus 3C pro Substrate Optimization Yields Efficient Substrates for Use in Peptide-Prodrug Therapy.
Miles, Linde A., et al. PloS One 10.6 (2015): e0129103.
The function of the milk-clotting enzymes bovine and camel chymosin studied by a fluorescence resonance energy transfer assay.
Jensen, Jesper Langholm, et al. Journal of Dairy Science 98.5 (2015): 2853-2860.
A comparison of modular PEG incorporation strategies for stabilization of peptide-siRNA nanocomplexes.
Lo, Justin H., et al. Bioconjugate Chemistry (2016).
Dilraj Lama, et al. Molecular dynamics simulations of pro-apoptotic BH3 peptide helices in aqueous medium: relationship between helix stability and their binding affinities to the anti-apoptotic protein Bcl-X(L). J Comput Aided Mol Des. 2011 May;25(5):413-26. : https://pubmed.ncbi.nlm.nih.gov/21523491/





